A History of DNA Sequencing and How It Has Changed the Way We Do Science at LSU
April 21, 2026
It’s 1979. Imagine carrying a 60-pound liquid nitrogen tank up a mountain in the Andes, on the off chance that you might sight a bird new to science. You are doing this to preserve a tissue sample from a specimen you might capture, but not for yourself. You are doing this so museum scientists can study your bird's genetics for decades to come. No serious technology for analyzing DNA even exists … yet.
Fast forward to 2026. The LSU Museum of Natural Sciences holds one of the largest collections of vertebrate genetic resources in the world, with over 200,000 samples. These samples come from specimens of birds, mammals, herps (reptiles and amphibians), and fish collected all over the world, from the U.S. to Southeast Asia to Papua New Guinea.
An artistic representation of DNA.
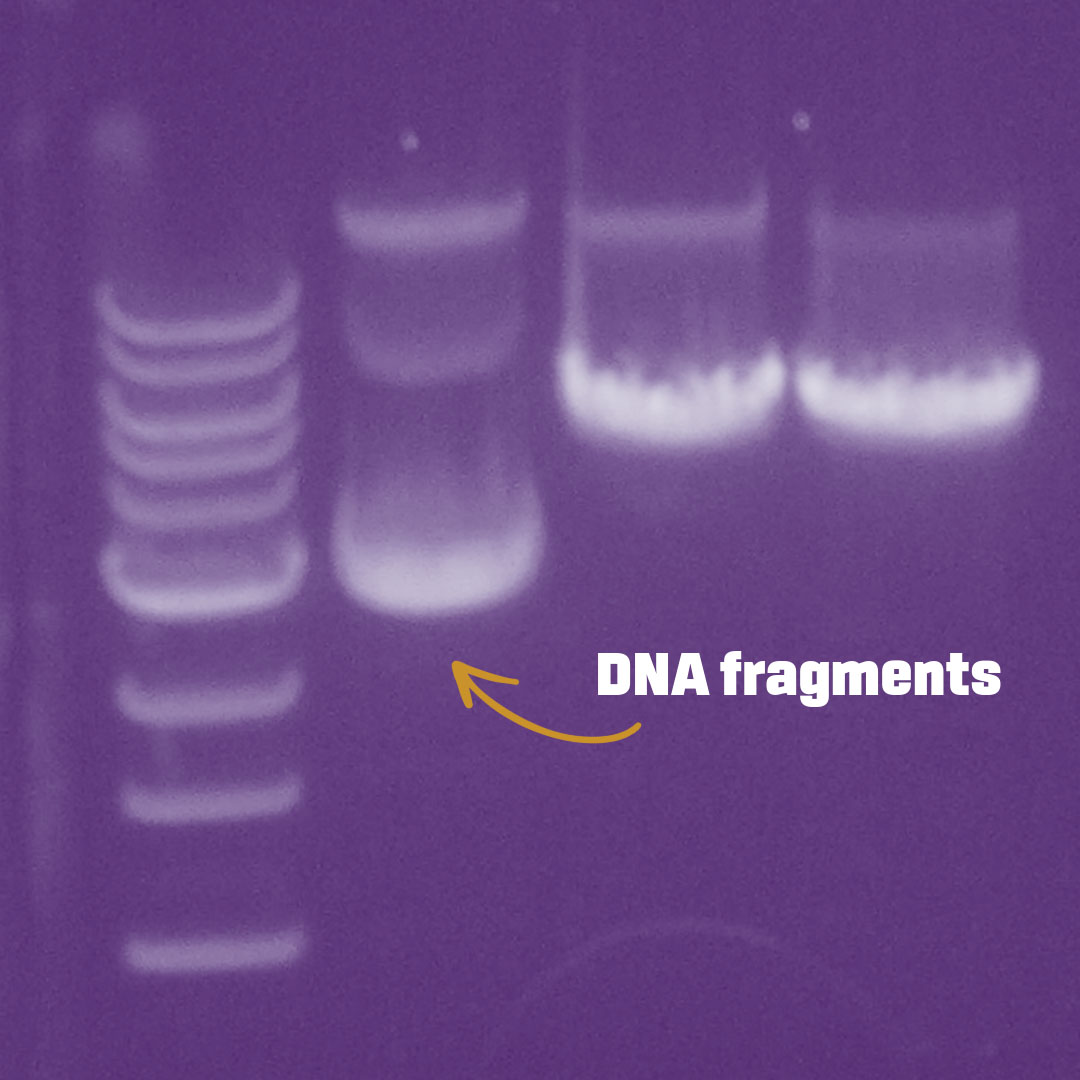
Gel electrophoresis image of DNA fragments
Many of these samples were collected and stored in liquid nitrogen – a condition critical to preserving sensitive molecules – before the technologies even came along to analyze most of their molecular signatures.
“The collection was officially created in 1979,” said Gregory Thom, curator of genetic resources at the museum. At first, researchers created private collections of tissue samples from field expeditions, but quickly began combining them.
In the 1970s, rudimentary techniques for analyzing DNA – lovingly referred to as the blueprint of life, the molecule that holds the instructions for your body to function – were just coming online. At first, museum scientists could only look for differences in DNA code indirectly. If they wanted to see how human DNA differed from mouse DNA, they used enzymes, or “molecular scissors,” to cut the DNA at specific sites. The resulting DNA fragments would vary in size and cut patterns (like paper snowflakes) across organisms and species.
Then came two key developments. First came a method for capturing, letter by letter, the code of a growing DNA strand by tracking the addition of labeled nucleotides (the “letters”). Then came the Polymerase Chain Reaction (PCR) technique, invented in 1985 by Kary B. Mullis. With PCR, scientists could make millions of copies of a small amount of DNA present in a sample.
Fun Fact: PCR uses a protein isolated from a heat-loving bacterium that can build new DNA strands from a template. It works at high temperatures, enabling an efficient heat-cycling process that copies DNA. Experts who regularly sequence DNA for research, including Camille Cannon, Genomics Research Specialist at LSU Health Shreveport, continue to use PCR today, every day.
Suddenly, researchers could efficiently amplify and then decode a specific region of DNA they were interested in. Museum scientists at LSU often examined a segment of DNA within mitochondria that steadily changes over evolutionary time; it is useful for distinguishing animal species from one another, and it was the first step towards building the tree of life.
“Our museum researchers at LSU were really pioneers,” Thom said. “I don't think they could even imagine that 40 to 50 years later, we would be sequencing whole genomes left and right in a very straightforward way.”
LSU researchers made the preservation of high-quality tissues from field-collected specimens a standard practice. They knew the technology would come. They stored tissue samples in liquid nitrogen, much colder than the standard laboratory -80°C freezer, to better preserve DNA, RNA, and proteins for yet-to-be-invented analysis techniques.
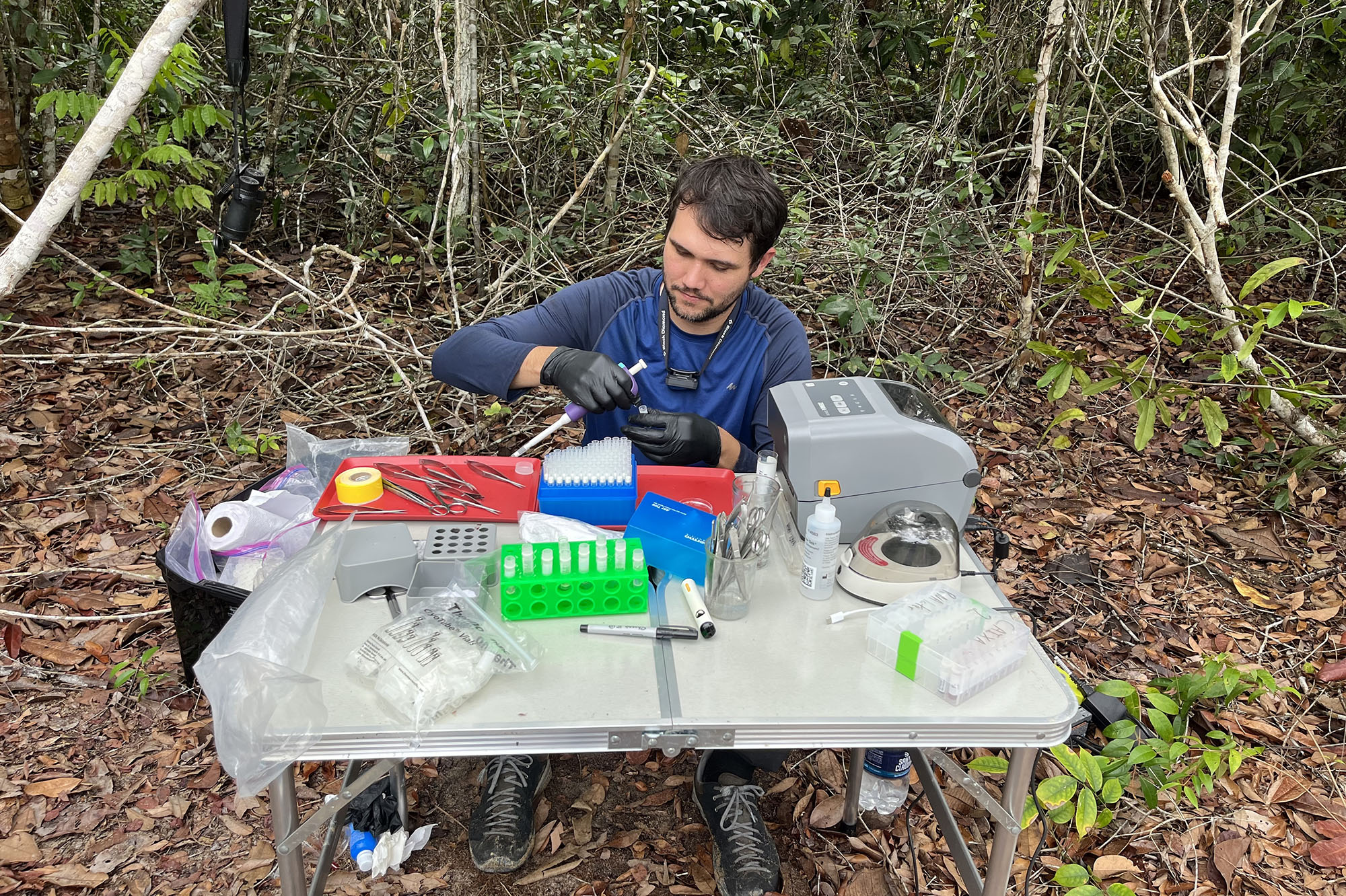

Gregory Thom from the LSU Museum of Natural Sciences processing samples for DNA analysis in the field.
Same Samples, Ever More New Data
Preserving as much data as possible from a specimen, even if that data isn’t accessible to researchers at the time of collection, is one way museum experts ensure no animal is captured and euthanized without a purpose. A single specimen can provide data for countless research projects and conservation efforts for years to come.
“Hundreds, if not thousands, of research papers were published using data from our museum samples in that PCR era, from the 1980s to the early 2000s,” Thom said.
Researchers soon began combining the molecular “scissor” technique with new, faster techniques for decoding DNA’s letter sequence. For faster sequencing, researchers would digest a genome – an organism’s entire DNA sequence – into short, single-stranded DNA fragments. They would then add in nucleotide building blocks (A, T, G, C), each labeled with a different dye, before using PCR to grow matching strands (DNA typically exists as a double helix with two paired strands). They would watch for color patterns at each PCR step as hundreds of matching DNA strands were synthesized letter by letter, ultimately decoding the sequence.
Of course, this required lots of analysis and computation to piece the fragmented code back together. Today, artificial intelligence is instrumental in matching these pieces together more quickly and accurately.
But early DNA sequencing was slow, labor-intensive, and costly, so it wasn’t usually feasible to decode an entire genome. Adding the molecular scissor technique allowed researchers to target only select areas of the genome they were interested in. But sequencing even 1-2% of the genome allowed museum researchers to begin building phylogenetic, or family, trees for animals.
Soon, however, the floodgates would open. Sequencing would become cheap and fast enough to enable routine whole-genome sequencing. But the raw data were already there, thanks to forward-looking LSU researchers.
“Around 2020, things really started changing towards whole genome sequencing,” Thom said. “Now, the sequencing technologies are so advanced, the protocols are so efficient, that we can sequence high-quality whole genomes for non-model organisms for fairly cheap.”
Whole Genome Sequencing Provides a Bigger Picture
Within the last decade, next-generation sequencing technologies have made rapid whole-genome sequencing accessible to researchers studying everything from evolution to cancer. Whole genome sequencing is faster and cheaper than ever, thanks to technologies that can analyze millions of DNA fragments simultaneously, often with the fragments attached to a chip or a flow cell surface.
“The first whole genome sequencing project, the Human Genome Project, cost billions of dollars. Today, we can sequence a whole genome for $250,” said Jovanny Zabaleta, a professor with LSU Health New Orleans and researcher with the LSU LCMC Health Cancer Center.
“Cancer is a really complex disease, an accumulation of mutations. It’s like a diabolical symphony. Just one instrument doesn’t make the symphony…. You have to put all the instruments together to make the sound of cancer.”
Jovanny Zabaleta, LSU Health New Orleans professor and LSU LCMC Health Cancer Center researcher
Whole genome sequencing involves looking at the entire DNA sequence of an organism, not just discrete sections. New technologies, including long-read DNA sequencing, are also revealing the DNA letter code in longer segments at a time. This is revealing parts of the genome – such as long repeating sections, complex insertions and deletions, and segments that may have jumped or moved around – that were difficult to piece together before. In these newly visible regions, researchers are finding new DNA mutations and variants associated with diseases like cancer.
Zabaleta is interested in understanding how different types of genetic variants, which vary in frequency across ancestries, relate to cancer risk and treatment response. Zabaleta’s lab at the LSU LCMC Health Cancer Center is currently sequencing genomes and transcriptomes from people across Louisiana, as well as surrounding areas and South America.
But what is the value of sequencing the whole genome, decoding every nucleotide letter base pair across every human chromosome, for cancer research? Why not just look for specific mutations or variants (small alterations in the DNA sequence) linked to cancer, such as BRCA gene mutations?
“Cancer is a really complex disease, an accumulation of mutations. It’s like a diabolical symphony,” Zabaleta said. “Just one instrument doesn’t make the symphony… You have to put all the instruments together to make the sound of cancer.”
Cancer emerges from a cascade of interacting factors. Not only are many of the genetic variants linked to cancer across populations unknown, but what really matters is how those variants interact with other genes, lifestyles, and environmental exposures. Those interactions and how they differ across populations are an underexplored frontier. Whole genome sequencing is the first step in putting all the pieces together.
“Here in Louisiana, we are a gumbo of ancestries,” Zabaleta said. “We have African American, Hispanic, Native American, Cajun, and Creole populations. Our DNA is a melting pot. How does having different mixtures of genetic variants based on your ancestry affect your susceptibility to cancer?”
Different ancestral populations are known to have unique genetic variants, varying frequencies of common genetic variants, and different cancer rates. But most of the tests used for clinical decision-making based on genetic risk factors come from research conducted in white European populations. That leaves many gaps in our view of minority populations' genomes.
“We need to include minorities in genetic sequencing analyses to understand what’s going on and how new arrangements in DNA influence disease risk or even your response to a particular drug,” Zabaleta said. “For example, we know that Hispanics respond differently to the same leukemia cancer drug treatments that white European-ancestry patients receive. And that's probably because of genetic differences that influence how these populations metabolize drugs. All of these things have to be taken into account when treating patients.”
Without sequencing entire genomes, you can’t fully visualize the combinations of different genetic variants that may significantly change cancer risk and optimal drug treatment plans.
“Cancer is not a single thing gone wrong; it’s an environment,” Zabaleta said. “It's a whole bunch of things that are going wrong at the same time.”
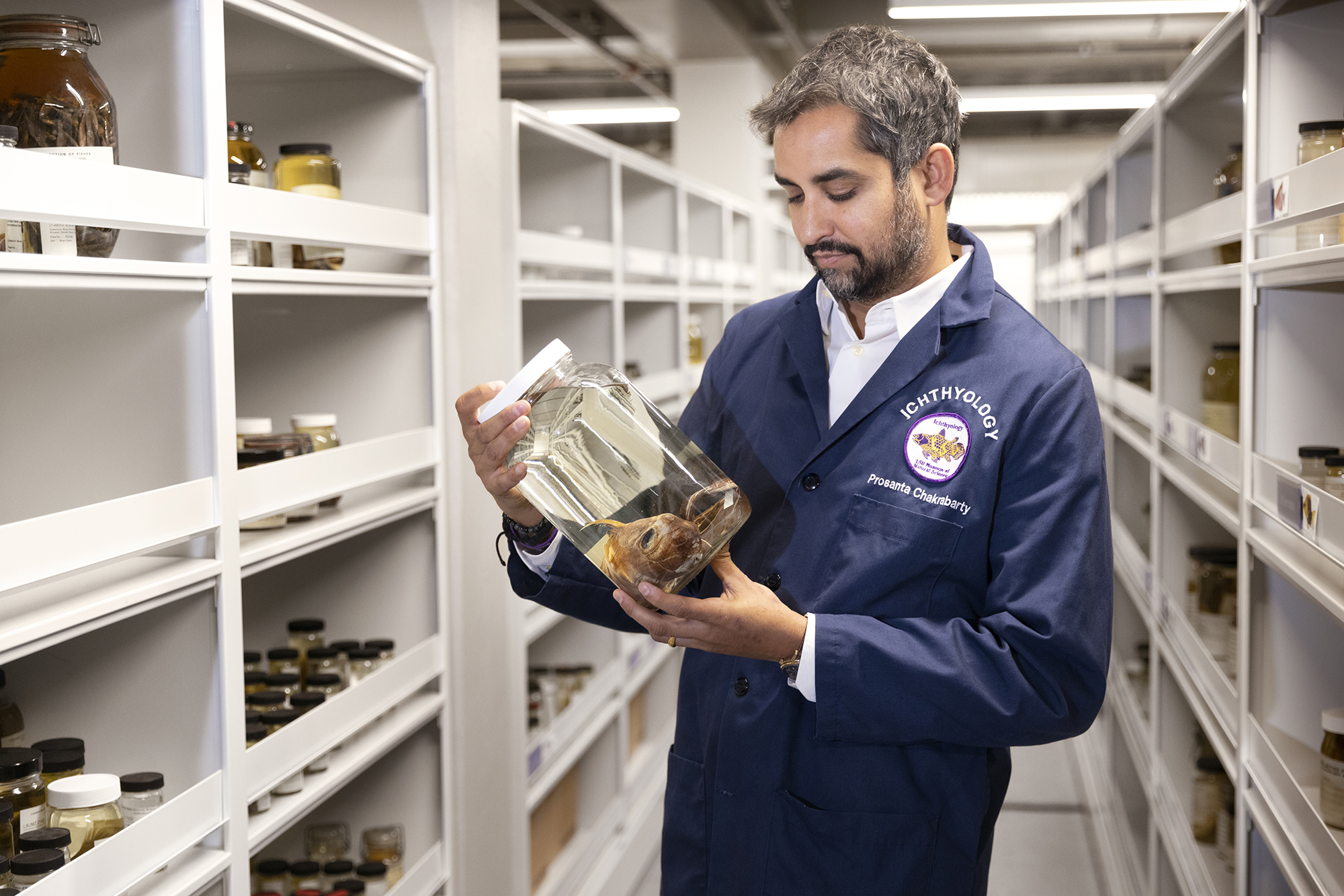

Prosanta Chakrabarty with a deep-sea fish in the LSU Museum of Natural Sciences.
From Cancer to Fishes: A Personal Tale of DNA
“It’s incredible to me, the level of personalized cancer care we can now do because we have the sequence of our entire DNA. And at the same time, we are also in a golden era of animal, plant, and fungal genetics and the ability to understand the tree of life.”
Prosanta Chakrabarty, professor and curator of fishes at the LSU Museum of Natural Sciences
Prosanta Chakrabarty, a professor and curator of fishes at the LSU Museum of Natural Sciences, has both a scientific and a very personal interest in genetics and how it is changing the way we approach science and healthcare.
In 2024, Chakrabarty was diagnosed with colon cancer.
“For a long time, the only option was chemotherapy, which I did 24 times. It sucks. It was worse than how the cancer made me feel,” Chakrabarty said.
But as modern genetic analysis techniques and molecular understanding of cancer improve, treatment options are improving as well.
“Now, doctors can analyze your genotype (genetic makeup) versus the genotype of the cancer,” Chakrabarty said. “They can analyze the genetic mutations present only in the cancer tissue and tailor treatment based on that.”
By analyzing exactly how cancer cells in his body differ genetically from his healthy cells, doctors can train Chakrabarty’s immune cells to recognize the downstream protein products of those genetic changes and attack cancer cells. This is a technique called immunotherapy.
“It’s incredible to me, the level of personalized cancer care we can now do because we have the sequence of our entire DNA,” Chakrabarty said. “And at the same time, we are also in a golden era of animal, plant, and fungal genetics and the ability to understand the tree of life.”
Fun Fact from Chakrabarty: Humans don’t have the largest genome in the animal kingdom. That record goes to an organism you’d least expect: the lungfish, an ancient-looking fish that can breathe air. In fact, LSU researcher Igor Schnieder and his lab have been exploring exactly why the lungfish genome is so large. (Hint: it seems lungfishes developed many copies of genes related to fin and limb development and regeneration.) Other organisms with massive genomes include a single-celled amoeba, rice, and axolotls.
“What makes something a living thing is a lot more complicated than we thought before we understood DNA, especially whole genomes,” Chakrabarty said.
On the other hand, birds have smaller genomes, not because they are less complex, but because they need small and efficient genomes so that their cells are light to help them fly! You don’t think of DNA as being something that takes up mass and weight; it’s invisible to the naked eye. But if you took all the DNA in your body and pulled it out into a single, linear strand, it could stretch beyond the Moon.
But how is genome size not directly related to complexity? Birds with small genomes certainly seem more complex than single-cell amoebas with huge ones.
There are many layers of complexity on top of the genome. These additional layers of control dictate what downstream products, including the RNAs and proteins, are produced (called gene expression) from the underlying genetic blueprint. The same gene in two organisms, or even duplications of that gene in the same organism, can produce very different downstream products.
This is where the power of the LSU museum’s liquid nitrogen tissue storage really comes into play. Flash-freezing samples preserves not just DNA, but also downstream RNA and protein products. Scientists around the world come to the museum to gain access to the data stored in these samples.
For example, LSU researcher Nick Mason is analyzing the protein products of genes involved in oxygen metabolism to understand how birds that live at high versus low altitudes use these genes differently. That wouldn’t be possible without flash-frozen tissue samples from birds living in different locations.
Studying animal evolution through genetics is not just an exercise in understanding these animals. It also helps us better understand ourselves. Chakrabarty is studying genes and gene mutations in fish, including a Hox gene mutation that may be related to hip dysplasia in humans. Other researchers look at animals with unique abilities, such as the ability to evade cancer or regenerate limbs, to understand how we might harness related genes and gene products to fight cancer and improve wound healing.
“We tend to think that understanding DNA unravels all the mysteries of who we are,” Chakrabarty said. “But we’re still at the early stages of figuring that out. It’s like the moon missions – we have to continue going to the moon because we’ve only just scratched the surface of the mysteries it holds.”
The Tale of the Army Ant Birds: The Power of Long-Read Sequencing
For decades, museum researchers relied on small genetic fragments and analysis of a handful of genes to build evolutionary trees. But much of the story of how various species evolved and what powered their incredible adaptations remained hidden.
“Before whole genome sequencing, we couldn't tell the full picture of what was going on with a particular species because we were restricted to seeing a very small part of their genome,” said LSU evolutionary biologist Gregory Thom. “Now, whole genomes are a kind of Pandora's box.”
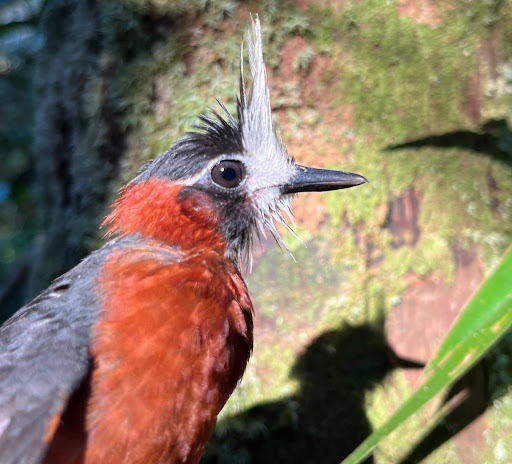

A species of bird that tracks army ants.
Thom’s lab is using whole-genome and long-read DNA sequencing to better understand one particularly complex ecological system: the relationship between army ants and the birds that follow them through tropical forests.
Army ants are top predators in tropical ecosystems, moving through forests in large, nomadic colonies. “They build a nest with their own bodies to protect the queen. And when it's time to move, they just move,” Thom said. “They prey on everything.”
Injured insects and animals that army ants feed on can also become dinner for bird thieves. Today, many bird species track and trail army ants to feed on the ants’ victims. Some species of birds are aggressive in stealing food from the ant swarm, while others hang back and feed on the leftovers. These birds don’t only face fierce competition from one another, but they also have to survive toxic and aggressive army ants.
Thom studies how different species of army ant-tracking birds have genetically adapted to this lifestyle. But it’s been difficult to do that until third-generation sequencing technologies like long-read DNA sequencing emerged.
For example, how do these birds track army ants in the first place? Studying this question led Thom’s lab to look at the evolution and genetics of olfaction.
“For a long time, people thought that birds couldn't smell, or at least they weren't very good at smelling,” Thom said.
That may have been because regions of the genome associated with olfaction in birds tend to be highly repetitive and unusually coiled, making them difficult to sequence with traditional techniques that fragment the genome. But long-read sequencing is revealing avian olfaction gene families in new detail.
“We're just figuring out that olfaction is something really important for some of these species,” Thom said.
Long-read sequencing is also allowing Thom to sequence other areas of bird genomes that have long been a nightmare to piece together, including genes associated with the immune system.
“We're also finding that there are major changes in the immune system of these species that track army ants, because they're exposed to a lot of pathogens as they congregate in large numbers around the ant swarms,” Thom said. “We are also studying the genetic basis of the blue skin some of the birds have developed around their eyes to symbolize dominance. This is something we couldn’t do even five years ago.”
With next-generation sequencing tools, the army ant bird system has become a living laboratory for understanding evolution and tracking down specific genes to see how other animals, including humans, might deal with toxic substances and pathogens.
Meanwhile, other genetic experts are moving beyond DNA analysis to explore downstream products – RNA and proteins – to better understand health and disease.
Jovanny Zabaleta with a student in the lab.
Beyond the Blueprint: RNA Sequencing for Cancer Research
While the ability to sequence entire genomes quickly has brought a lot of excitement and hope to fields like cancer research, it has also raised many more questions, revealing just how complex the genetic control of health and disease really is.
“You can have a mutation in your DNA, but only under certain conditions will that mutation actually lead to cancer,” Zabaleta said. “Cancer is not only genetics. It's also where you live, what you eat, what you do, what you breathe.”
Imagine a first draft of a building blueprint. It holds all the plans for the building. But in practice, buildings often differ from the original plans due to unforeseen site conditions, material constraints, and any number of issues that arise that require modifications. Our cells are masters at translating their DNA blueprints in different ways to suit different roles, environments, and constraints.
Diseases that are directly and inevitably caused by single genetic mutations are rare. Cancer typically results from multiple mutations interacting with one another, other genetic variants, and lifestyle and environmental factors. To understand how any given mutation contributes to the development of cancer, researchers like Zabaleta have to follow its downstream effects on biology across multiple steps, from blueprint to final construction.
“What is the transcript, the RNA that is being generated with that mutation?” Zabaleta said. “And what is that RNA modifying? What proteins are being produced from it, and how do they differ from the non-mutated proteins?”
In the end, proteins are the workers of cellular biology, and most pharmaceutical drugs are designed to target them. “So we're basically working backward, trying to find from the genome what might happen going forward,” Zabaleta said. “If you put all those things together, that's going to be a much bigger and precise picture of what's actually going on in the cell.”
New technologies are enabling Zabaleta to sequence RNA transcripts from patients’ cancer cells, identify which RNAs and proteins are being made in cancerous and non-cancerous cells, and even create maps of these “expression” patterns within tissues.
“We can follow patients over time and see how cancer cells, for example, change noncancerous cells around a tumor, changing their expressed RNA and proteins,” Zabaleta said.
These and other studies are leading to a finer-grained understanding of how cancer develops, how it evades the immune system, and how it reacts to different drugs.
Zabaleta’s lab, working with Dr. Lucio Miele’s lab at the LSU Health New Orleans Cancer Center and several other researchers at LSU Health New Orleans, recently published a research paper for which they sequenced the genomes of over 250 Louisiana patients with triple-negative breast cancer. They layered together different datasets, from DNA analyses to RNA analyses to maps of cells’ expressed RNA patterns within biopsied tissue. By doing this, the researchers can better understand what is happening within breast cancer tissues and which patients, based on observed patterns, might respond best to specific treatments.
“I'm really blessed that we can do that here at the Cancer Center of LSU Health New Orleans, because we have all the tools needed to do that,” Zabaleta said.

Camille Cannon in the Genomics Core at LSU Health Shreveport.
Supporting the Research Enterprise with Sequencing Expertise
Genetic and other “omic” sequencing analyses (looking at RNA, proteins, and metabolites) are increasingly a mainstay in labs studying biology, health, and disease. Access to the resources and training needed to fully leverage multi-omic analyses is now critical. Campuses across the LSU system are rising to the challenge by creating omics core facilities.
“Anything involving DNA or RNA, I’ve probably done it, including single-cell sequencing,” said Camille Cannon, a research specialist at the Genomics Core at LSU Health Shreveport. “We get samples all across Louisiana. I work with researchers at different LSU campuses and at Tulane.”
Cannon is increasingly helping researchers with single-cell DNA and RNA sequencing. This involves isolating single cells within a tissue, such as a single cancer cell, to analyze their genomes and gene expression patterns.
“RNA is the message transcribed from your DNA. ... With single-cell sequencing, we can map out and know what types of cells are expressing what, which is pretty cool.”
Camille Cannon, research specialist at the Genomics Core at LSU Health Shreveport
“RNA is the message transcribed from your DNA. If I have a professor looking at a cancer tissue sample, for example, and we just chop up the tissue and isolate all of the RNA from that, we can’t tell which RNA transcripts are coming from which types of cells in the tissue,” Cannon said. “With single-cell sequencing, we can map out and know what types of cells are expressing what, which is pretty cool.”
Cannon is working with researchers at LSU Health Shreveport, including Omar Franco-Coronel, to look at gene expression in single cancer cells.
“There is a lot of interest in cancer research to look at RNA expression and link that back to DNA mutations,” Cannon said. “Does this DNA mutation directly increase a particular RNA transcript, or is it causing some other thing to happen in the cell that's ultimately affecting that RNA expression?”
Cancer researchers are increasingly looking at molecules known as microRNAs, because they can bind to other RNA molecules and increase or decrease their expression into protein products.
“Pharmaceutical companies are interested in microRNAs now, because they are finding that they might be able to target some of these microRNAs to control gene activity in cancer cells,” Cannon said.
As applications for increasingly refined DNA and RNA sequencing expand, core facilities and experts in both traditional and out-of-the-box sequencing methods, like Cannon, can help researchers answer evolving research questions.
Learn More: DNA Day Spotlight: Supporting Discovery Through Genomics at LSU Health Shreveport
Fun Fact from Camille Cannon:
Most of our DNA is actually “junk” DNA, or regions that don’t code for proteins. That doesn’t mean it is useless; new sequencing technologies are teaching us more and more about the hidden functions of these genetic regions.
For example, they often contain mobile elements, sections known to jump around the genome. Many of these mobile elements are remnants of ancient viral infections. Researchers have classically used them to study evolutionary history, but they are also finding that some diseases arise when these elements jump into our more active DNA regions.
New Frontiers
DNA, RNA, and protein analysis techniques will continue to improve, revealing aspects of our genetic histories and mechanisms underlying health and disease that we didn’t know existed.
“There's just so much data now,” Chakrabarty said. “We may go into the jungle to collect animal species for our research, but when we come back to the lab, we have another jungle to explore within their genomes. The possible insights still hidden in these samples are mind-boggling.”
However new sequencing technologies evolve, one thing is for sure: LSU researchers will be there to use those tools to glean new insights into our world and improve human health.


